 
Lee J, Boutz DR, Chromikova V, Joyce MG, Vollmers C, Leung K, et al. Lineage structure of the human antibody repertoire in response to influenza vaccination. Jiang N, He J, Weinstein JA, Penland L, Sasaki S, He XS, et al. Defining antigen-specific plasmablast and memory B cell subsets in human blood after viral infection or vaccination. Data from other cell input are in Figure S6B in Supplementary Material.Įllebedy AH, Jackson KJ, Kissick HT, Nakaya HI, Davis CW, Roskin KM, et al. 
The overlapping clones were compared between two adjacent sub-samplings and overlap percentage was calculated by dividing the number of overlapping clones by the total number of clones observed in the deeper sub-sampling. (D) The percentage of overlapping clones with single RNA copy at different sequencing depths by sub-sampling in 20,000 naïve CD8 + T cells for three RNA input amounts. Data from other cell inputs are in Figure S6A in Supplementary Material. Sequencing reads were subsampled to different depth and unique CDR3s were tallied. (C) Rarefaction curve of number of unique CDR3s with single RNA copy in 20,000 naïve CD8 + T cells for three RNA input amounts. (B) Comparison of rarefaction curve of detected RNA molecules and unique complementarity-determining regions 3 (CDR3s) in 20,000 naïve CD8 + T cells for three RNA input amounts. Data from other cell inputs are in Figure S4 in Supplementary Material. (A) Rarefaction curve of detected TCR RNA molecules before and after error correction on molecular identifiers (MIDs) in 20,000 naïve CD8 + T cells for three RNA input amounts. MID Clustering-based IR-Seq is capable of accurate digital counting of T cell receptor (TCR) RNA molecules. Short vertical black lines indicate nucleotide differences between two TCR sequences. Top: without sub-clustering, chimera sequences are generated when different TCR RNA molecules are tagged with the same MID bottom: TCR RNA molecules that are tagged with same MID are sub-clustered to reveal truly represented TCR sequences. (D) Illustration of consensus TCR sequence building without (top) and with (bottom) sub-clustering. Data from other cell inputs are in Figure S2 in Supplementary Material. Number of unique CDR3s in three libraries made with three different RNA inputs from sorted one million naïve CD8 + T cells are shown here. The theoretical percentage of MIDs with sub-clusters was calculated by Eq. 2 in Section “Materials and Methods.” (C) Rarefaction curve of unique complementarity-determining regions 3 (CDR3s) with or without sub-clustering. (B) The theoretical percentage of MIDs with sub-clusters is approximately linearly dependent on copies of target molecules when copies of target molecules are less than 5,000,000 (bottom right insert). 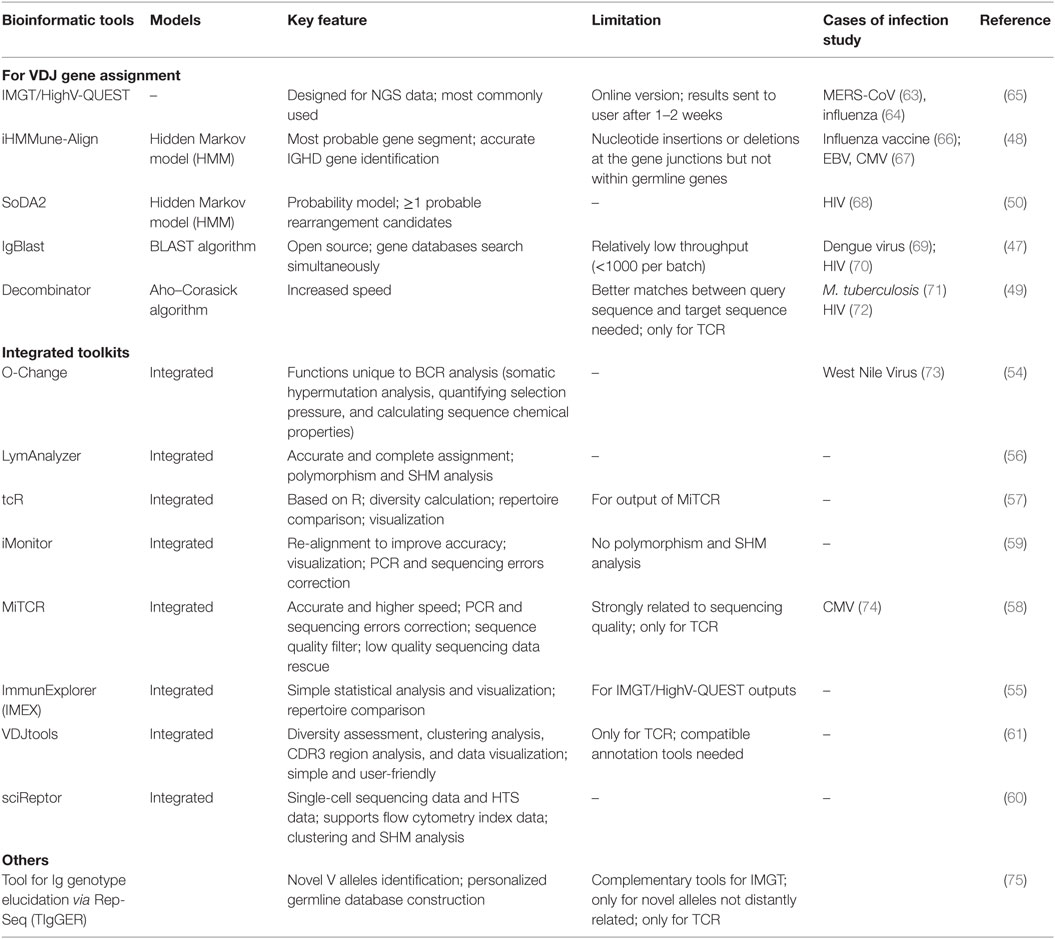
Line represents linear regression fit, F-test on the slope, p < 10 −9. (A) The percentage of observed molecular identifiers (MIDs) containing sub-clusters is linearly dependent on RNA input, which is defined as cell number multiplied by percentage of RNA (e.g., 20,000 cells with 10%RNA is equivalent to 2,000 RNA input). MID Clustering-based IR-Seq improves accuracy of T cell receptor (TCR) diversity estimation with sub-clustering. The demonstrated accuracy, sensitivity, and wide dynamic range of MIDCIRS TCR-seq provide foundations for future applications in both basic research and clinical settings.ĬMV-specific T cells MID clustering-based IR-Seq TCR repertoire sequencing molecular identifiers naïve T cells sub-clustering. Further, we showed that MIDCIRS enables a sensitive detection of a single cell in as many as one million naïve T cells and an accurate estimation of the degree of T cell clonal expression.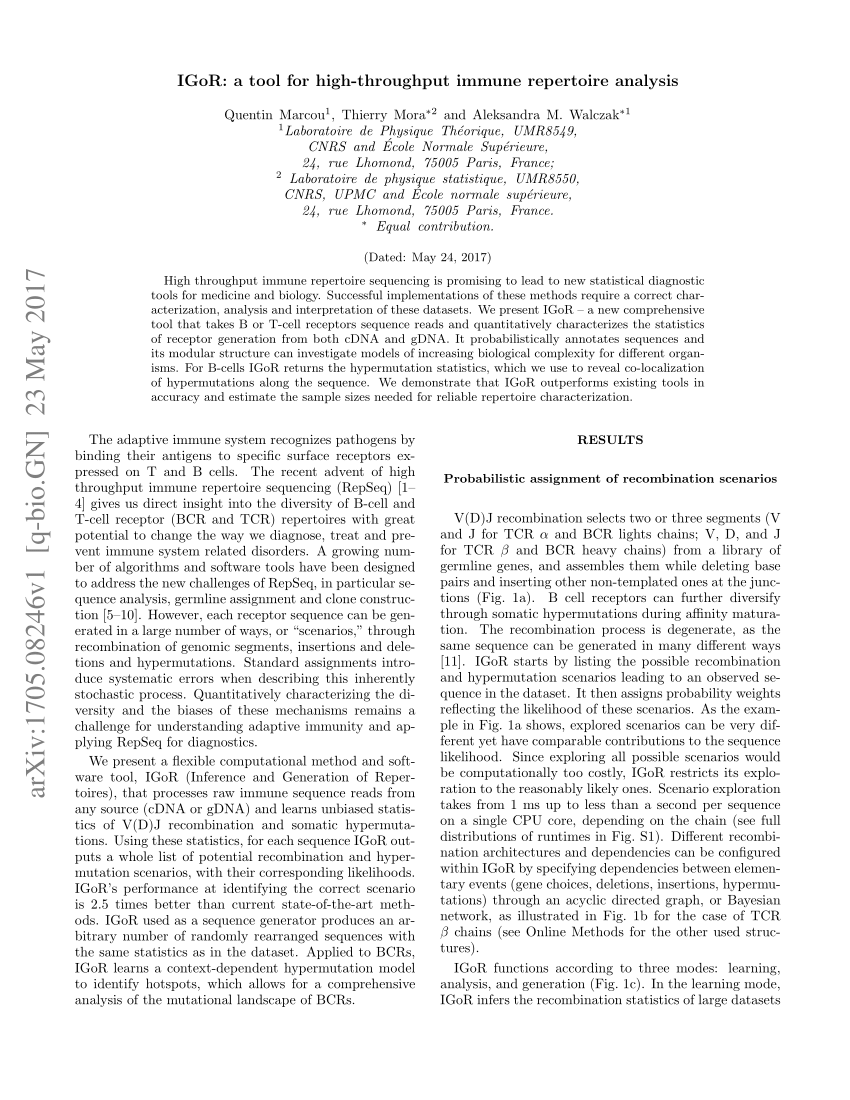 
First, we demonstrated the necessity of performing MID sub-clustering to eliminate erroneous sequences. Here, we describe using an MID Clustering-based IR-Seq (MIDCIRS) method to quantitatively study TCR RNA molecule copy number and clonality in T cells. This limited the application of TCR repertoire sequencing (TCR-seq) in clinical settings, such as detecting minimal residual disease in lymphoid malignancies after treatment, evaluating effectiveness of vaccination and assessing degree of infection. However, evaluating the sensitivity to detect rare T cells and the degree of clonal expansion in IR-seq has been difficult due to the lack of knowledge of T cell receptor (TCR) RNA molecule copy number and a generalized approach to estimate T cell clone size from TCR RNA molecule quantification. Unique molecular identifiers (MIDs) have been demonstrated to effectively improve immune repertoire sequencing (IR-seq) accuracy, especially to identify somatic hypermutations in antibody repertoire sequencing. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
